Announcement
NEW: Over 100 African researchers begin training... more
Latest data resources
December 8, 2022
Pf7, the newest version of our largest Plasmodium falciparum dataset, contains over 20,000 whole genome sequences. The paper describing the results was published in Wellcome Open Research in January 2023.
Started 2021
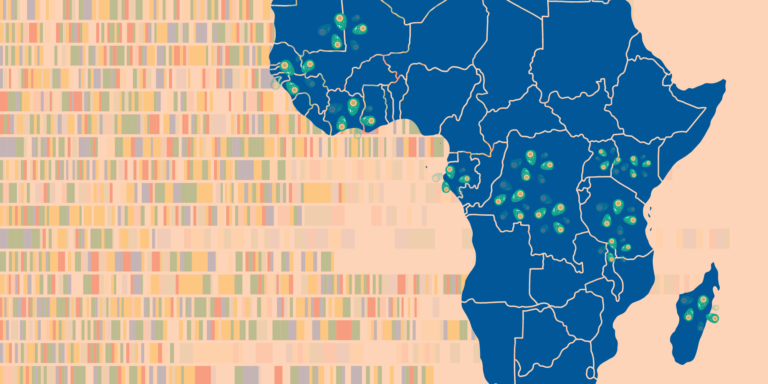
The MalariaGEN Vector Observatory has sequenced more than 15,000 samples in the Anopheles gambiae complex from 26 countries.
February 11, 2022
The Pv4 dataset contains genome variation data on 1,895 worldwide samples of Plasmodium vivax, the second-deadliest form of malaria.
News from around the network
News
Changes to MalariaGEN cloud data access
To help us continue providing cloud data resources for the community, we are making some changes to how data are accessed. If you are actively using MalariaGEN data, or are planning to use it in the future, please read this carefully and follow the steps below.