About this study
Malaria, caused by the parasite Plasmodium falciparum, remains the most important disease in sub-Saharan Africa and a leading cause of global morbidity and mortality. As part of an effort to characterise the Kassena-Nankana District (KND) of Northern Ghana for future vaccine trials, we set up a large, frequency-matched, case-control study of severe malaria.
A number of malaria candidate genes, and more recently genome-wide association studies, have identified genomic loci associated with severe P. falciparum malaria disease. Notable among these is the protection conferred by the sickle-cell locus (HbS) in the beta-globin gene (Jallow M et al, 2009). Though Mackinnon and colleagues in Kenya estimated that host genetic factors contribute about a quarter of total malaria protection, only 2% of the total can be attributed to HbS (Mackinnon MJ et al, 2005), the ability of these genotypes to censor severe disease remains a strong component of the molecular mechanisms underpinning host response to malaria disease.
Summary
A frequency-matched and an individually-matched case-control study were conducted in the Kassena-Nankana District between 2002/04, and 2007/08, respectively, with the aim of identifying human genetic determinants of resistance to severe malaria in Northern Ghana.
Severe malaria cases were recruited from the paediatric ward of the Navrongo War Memorial Hospital (NWMH), which serves the health needs of the Kassena-Nankana District population. Cases consisted of children (aged 6-60 months) who were diagnosed with severe malaria (SM). Criteria for diagnosis of SM were those included in the standard WHO definition. Cases were matched to controls on age category, sex, ethnicity and location. Two control categories were used; mild malaria and healthy (asymptomatic, community) controls. Two controls were recruited per case for each category of controls. The Navrongo Health and Demographic Surveillance System keeps vital records of the KND population and played a central role in the location and recruitment of community controls. Mild malaria controls were recruited from four out-patient clinics, including the out-patient department of the NWMH.
Clinical data and DNA samples were contributed to the MalariaGEN Consortial Project 1 (CP1) along with those of 10 other case-control studies from a total of 11 malaria-endemic countries. As part of the sample handling process, baseline genotyping data was generated for a number of malaria–associated single nucleotide polymorphisms (SNPs) and the appropriate data has been returned to each site for site-specific analysis. A total of 69 SNPs at candidate genes (selection based on previous reports of association with severe malaria or on their likely biological role in malaria infection/disease) will be included in our analysis.
Single- and multi-locus analysis will be conducted using various multivariate logistic regression models to assess the relationship between these polymorphisms and well defined clinical phenotypes. Phenotypes that could potentially be tested include severe malaria and sub-phenotypes such as severe malarial anaemia, hyperlactatemia and respiratory distress.
Study site description
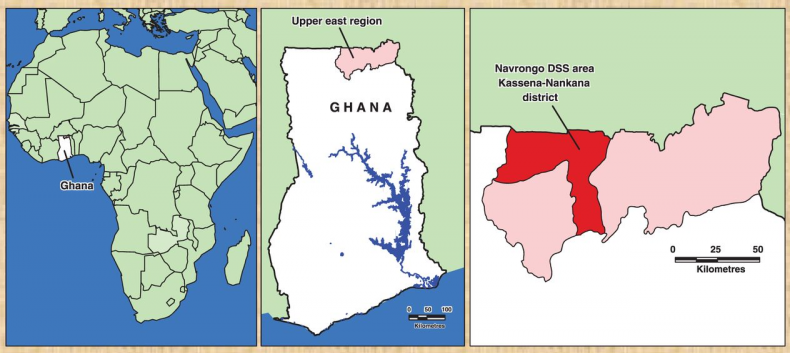
The studies were conducted in the Kassena-Nankana District of the Upper East region of Northern Ghana. The district covers an area of 1675Km2 of Sahelian savannah along the Ghana-Burkina Faso border. The population is estimated at 152,000 inhabitants, the majority of whom are engaged in subsistence farming. The major ethnic groups in the area are the Kassem and the Nankam with a minority of Buli. Most people live in traditional family housing units called compounds that typically house multi-families (different households) of common ancestry.
Annual rainfall averages 850mm, almost all of which occurs in the wet months of July – November with the rest of the year being relatively dry. One of the main features of the area is a large reservoir in the district, which provides water for irrigation throughout the year. Malaria epidemiology has been extensively characterized in this part of Northern Ghana (Owusu-Agyei S et al, 2001; Owusu-Agyei S et al, 2002; Koram KA et al, 2000; Koram KA et al, 2003; Oduro AR et al, 2007). Malaria transmission in the KND is stable but with a marked seasonality in intensity that coincides with the months of rainfall. The overall entomological inoculation rate at the time of these studies was estimated to be >400 infective bites/person-year (Appawu M et al, 2004), but current estimates are around 150 infective bites/person-year. The main malaria vectors are Anopheles gambiae s.l and A. funestus, constituting about 94.3% of the vector population.
One hospital, the Navrongo War Memorial Hospital (NWMH) serves as a referral facility for the KND residence while about six public Health Centres are strategically located across the districts to support the NWMH. In addition, most communities have a Community Health Planning Services (CHPS) compound that is run by community health officers and serves as the first line of contact for basic health services that includes door-to-door health services and early identification of referral cases.
The Navrongo Health Research Centre maintains the Navrongo Health and Demographic Surveillance System (NHDSS) in the KND. The NHDSS collects and updates vital records of the KND population and was a key resource for these studies. The NHDSS database was used to identify potential participants to be recruited as healthy controls based on the matching criteria.
Methods
A frequency-matched and an individually-matched case-control study were conducted in the Kassena-Nankana District between 2002/04, and 2007/09, respectively. Severe malaria cases were recruited from the paediatric ward of the Navrongo War Memorial Hospital (NWMH) which serves the Kassena-Nankana District.
Cases consisted of children (aged 6-60 months) who were diagnosed with severe malaria at the NWMH. Criteria for diagnosis of severe malaria followed the definitions stated by the World Health Organization i.e. the presence of asexual parasitaemia and at least one of the following: cerebral malaria (Blantyre coma score of 3 or below and coma persists for more than 30 minutes after fits have ceased); repeated or prolonged generalized convulsions; severe malarial anaemia (haemoglobin <5g/dl); respiratory distress (presence of alar flaring, intercostals or subcostal chest recession, use of accessory muscles of respiration, or abnormally deep respiration); hypoglycaemia (blood glucose <2.2mM/l); circulatory collapse (systolic blood pressure <50mmHg); renal failure (urine output less than 12 ml/kg/24 hours or serum creatine >3.0mg/dl); hyperpyrexia (anxillary temperature >39oC, hyperlactataemia (blood lactate >2.0mmol/l) and impaired consciousness). Further details may be reviewed (Oduro AR et al, 2007; Osafo-Addo AD et al, 2008).
Using the case as an index, the NHDSS was used to recruit two groups of population controls; mild malaria and community (‘healthy’) controls (a fraction of whom had asymptomatic parasitaemia). Two controls were recruited per case in each control category. These were matched by age category (6-24 months and 25-60 months), gender, location (areas of residence) and ethnicity prior to 2007, after which they were individually matched to severe malaria cases using the same criteria.
A standardised case report form (CRF) was created by MalariaGEN and used by all sites to collect standardised clinical data. The relevant data fields as per the CP1 CRF were extracted from CRFs of the studies conducted prior to MalariaGEN (ie 2002/04 study conducted in Ghana and all other sites). Data were uploaded onto the MalariaGEN central repository via secure web-based software developed by MalariaGEN. Here, the integrity of the data was checked and data was standardised and amalgamated.
Genomic DNA was extracted from whole blood, at the Noguchi Memorial Institute for Medical Research, using the Chelexmethod or Nucleon™ BACC2 Genomic DNA extraction kit® (Gen-Probe Life Sciences Ltd., Manchester, UK) using manufacturer’s instructions. Aliquots of the DNA samples were shipped to the MalariaGEN Resource Centre in Oxford for further processing and quality control for quantity, quality (by genotyping) and confirming appropriate clinical data was available. Baseline genotype data for 69 malaria-associated SNPs was generated for all contributing samples; briefly, samples underwent a primer-extension pre-amplification (PEP) step (Xu K et al, 1993; Zhang L et al, 1992) prior to genotyping on the Sequenom® MassArray® platform. Following curation, the genotype data were returned to the PIs for local analyses.
| Number | Gender: n (%) | Age in years: n (%) | Ethnicity: n (%) |
|---|---|---|---|
| Malaria cases: 2685 | Male: 1369 (51)
Female: 1059 (39) Not recorded: 263 (10) |
<1: 830 (31)
1-2: 1176 (44) 2-5: 677(25) Not recorded: 2(<1) |
Kasem: 1526 (57)
Nankam: 792 (30) Other: 360 (13) Not recorded: 6 (<1) |
| Healthy controls: 2378 | Male: 1126 (47)
Female: 957 (40) Not recorded: 301 (13) |
<1: 573 (24)
1-2: 872 (37) 2-5: 503 (21) Not recorded: 430 (18) |
Kasem: 1302 (54)
Kasem mixed: 103 (4) Namkam: 102 (4) Nankam: 706 (29) Other: 197 (8) |
Ethics
These studies were reviewed and approved by scientific and Institutional review boards of the Noguchi Memorial Institute for Medical research, the Navrongo Health Research Centre, The Ghana Ministry of Health Ethics review committee, and the U.S. Naval Medical Research Unit # 3 (proposal number: NMIMR-IRB CPN 016/01-02; ID NMIMR-IRB CPN 029/05-06).
At recruitment, written informed consent was obtained from participants (cases and controls) and witnessed by an individual who was not part of the research team. Individual informed consent was obtained from parents or legal guardians of children.
Additional contributors
- Abraham R Oduro, Navrongo Health Research Centre, Ghana
- Abraham VO Hodgson, Ghana Health Service, Ministry of Health, Ghana
- Francis Nkrumah, Noguchi Memorial Institute For Medical Research with Navrongo Health Research Centre, Ghana
- Frank Atuguba, Noguchi Memorial Institute For Medical Research with Navrongo Health Research Centre, Ghana
- Nana Akosua Ansah, Noguchi Memorial Institute For Medical Research with Navrongo Health Research Centre, Ghana
- Nathan Mensah, Noguchi Memorial Institute For Medical Research with Navrongo Health Research Centre, Ghana
- Patrick A Ansah, Noguchi Memorial Institute For Medical Research with Navrongo Health Research Centre, Ghana
- Thomas Anyorigiya, Noguchi Memorial Institute For Medical Research with Navrongo Health Research Centre, Ghana
- Victor Asoala, Noguchi Memorial Institute For Medical Research with Navrongo Health Research Centre, Ghana
- William O Rogers, Noguchi Memorial Institute For Medical Research with Navrongo Health Research Centre, Ghana
Acknowledgements
We acknowledge all our study participants, especially the parents or legal guardiansof the children who participated in these studies. Our gratitude goes to the people of the Kassena-Nankana districts for their support for our studies. Our appreciation and gratitude goes to the study team at the Noguchi Memorial Institute for Medical Research, especially Afrane Forson and Josephine Quagraine. Also, we wish to thank the study team and staff of the Navrongo Health Research Centre, particularly Raymond Aborigo, Daniel Tindanbil, Dickson Amugsi for their contributions.
These studies were supported by the US Naval Medical Research Centre and National Institute of Allergy and Infectious Diseases, National Institute of Health contract NO1 A195363 to the Noguchi Memorial Institute for Medical Research and subcontracted to the Navrongo Health Research Centre. Additional support for these studies was from a MalariaGEN Consortial sub-grant award, and additional downstream laboratory support by MalariaGEN.
Publications
-
Insights into malaria susceptibility using genome-wide data on 17,000 individuals from Africa, Asia and Oceania
MalariaGENNature Communications, 2019; 10 5732
-
Characterisation of the opposing effects of G6PD deficiency on cerebral malaria and severe malarial anaemia
Clarke, Rockett et al.eLife, 2017; 6 e15085
-
Admixture into and within sub-Saharan Africa
Busby et aleLife, 2016; 5:e15266
-
A novel locus of resistance to severe malaria in a region of ancient balancing selection
Malaria Genomic Epidemiology NetworkNature, 2015; 526(7572) 253-257
-
Reappraisal of known malaria resistance loci in a large multi-centre study
Rockett et al.Nature Genetics, 2014; 46(11) 1197-204
-
A global network for investigating the genomic epidemiology of malaria
Malaria Genomic Epidemiology NetworkNature, 2008; 456(7223) 732-7
