Tell us a little about your research background and how you moved into operational work with MalariaGEN?
I came into operational management of scientific projects from a research background. I really enjoyed working on projects with a global health focus and started as an undergraduate working on schistosomiasis. Following my Master's, I completed a PhD at the Liverpool School of Tropical Medicine, investigating insecticide resistance in the dengue vector Aedes aegypti in South East Asia and Latin America.
I went on to work as a lab-based scientist with management responsibilities for two longitudinal studies. This is where I started to move away from physical laboratory work and became more involved in operations and management. When the opportunity arose to work as a Scientific Project Manager with our MalariaGEN partners, I jumped at the chance. I am part of Dr Sonia Goncalves’ team and lead the in-country implementation of our amplicon sequencing protocols in our MalariaGEN partner labs. I understand the importance of a smooth operations system, so I can hopefully offer the right support for partners when it comes to implementing amplicon sequencing in their labs.
Why has in-country amplicon sequencing become such a priority for MalariaGEN?
Historically amplicon sequencing protocols have been implemented in our Sanger labs. MalariaGEN partners would routinely send samples that were run through laboratory pipelines, with resulting data returned to the partners. Once the scientific foundations and the underlying technology required for genomic surveillance was developed, the ambition was to decentralise the model and translate it into practical tools and protocols that could help national malaria control programmes (NMCPs).
From a malaria control perspective, it’s essential that NMCPs have tools to monitor resistance so they know what drugs and insecticide to use to control malaria. Surveillance also provides an early warning of whether the interventions are causing resistance to rise and whether they need to change strategy.
Why has one of the regions of focus for the development of an amplicon sequencing toolkit been West Africa?
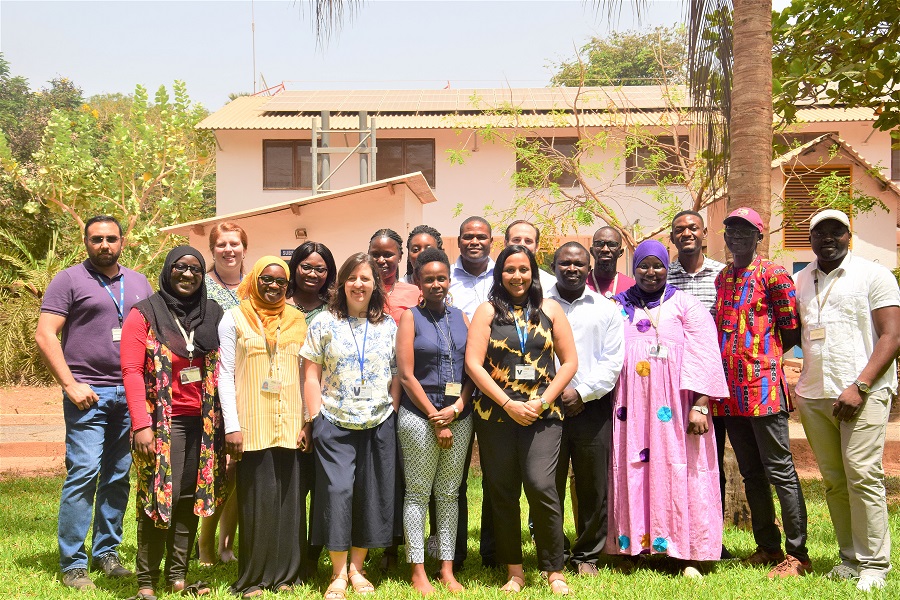
Focusing on West Africa allowed us to optimise surveillance tools for different use cases in a range of epidemiological settings. Many areas of West Africa have a very high transmission rate, for example in northern Ghana, but you have areas of low transmission, for instance the coastal region of The Gambia and north Senegal. Operationally developing two different labs in the same region helps develop critical mass and regional cooperation. It was also very important to work with partners who had strong engagement with NMCPs for the success of this project.
From an implementation standpoint, our partners at the University of Ghana and MRC Unit The Gambia at LSHTM had access to excellent lab infrastructure and sequencing equipment. They had outstanding leadership and institutional support, renowned malaria experts and among the most accomplished technical researchers working in the field. They also had significant experience in epidemiological, lab and analytical aspects of malaria research.
What knowledge and technology transfer were required to build operational systems in-country?
The first thing we had to do was establish that the protocols we had developed were suitable to use in an external lab. At the start it was a bit of trial and error and working with feedback from partners. But slowly and surely, we have built a set of protocols that are more suitable for labs in lower middle-income countries (LMIC). From an operational perspective you have some building blocks to set up an amplicon operation in country, but it is not a one size fits all approach and we work very closely with our partners to tailor implementation for an individual lab.
It is also worth bearing in mind when working with external funders, you would also need to arrange research collaboration agreements, due diligence around good financial grant practise, and you need to work with teams across partner institutions to access the committed funding. From our experience these processes can take time. It is important to assess each partner’s overall set-up and identify any key resources required, be it equipment or staff, what their training requirements would be and establishing what supply chains are in place to facilitate the smooth running of the whole process.
What impact did the pandemic have on implementing amplicon sequencing in West Africa?
Like the UK, much of the world including The Gambia and Ghana went into national lockdown and this impacted on both the sample collections and laboratory processing of samples, which must have been so frustrating for partners who’d almost got everything up and running. However, they were amazingly resilient and worked with in country public health agencies so that samples were collected in parallel for both COVID-19 and malaria. It also had a huge impact on laboratory supply chains.
In terms of my role, I was used to remote working, but the pandemic meant I was not able to meet colleagues at Sanger on a day to day basis and pop into someone’s office with a quick question. We also couldn’t visit the partner labs and carry out hands-on training. Everything had to move onto an online platform and Zoom calls – but nothing can replace hands-on and face to face training, and I have missed visiting our partners, but we have all had to learn to adapt.
What has been the highlight of your work in the past year?
I think there have been several highlights. We are currently working with six MalariaGEN partners across West Africa and South East Asia. One of the amazing moments has been when our partners in Vietnam, The Gambia, Ghana and Indonesia completed their first runs. Whilst we still have a bit of optimising to do, this was an incredible next step. Another highlight was the SARS-CoV-2 sequencing that was led by our team in Ghana. This work has been published and their team’s achievements have quite rightly been recognised.
I think the COVID-19 pandemic has brought genomic surveillance into the spotlight. To successfully implement genomic surveillance into a routine surveillance programme you need to have buy-in from all stakeholders. With the pandemic the importance of genomics in surveillance has been highlighted and even the most sceptical amongst us saw the crucial role it played globally to support decision-making for disease control. Hopefully, moving forward, this will facilitate integrating genomic surveillance into routine malaria control operations in West Africa and beyond.