Shortly before his death in April 2023, Professor Dominic Kwiatkowski completed the final draft of a manuscript he had been working on for years. Now posthumously published, this sole-authored scientific paper represents a career of astonishing diversity and depth, ranging from genomics and bioinformatics, to epidemiology and public health.
Perhaps most significantly, Dominic’s final paper plots a course for the future, detailing the territory he wanted malaria genomics to explore next. Although the field will no longer benefit from Dominic’s direct leadership, colleagues are able to refer back to his thinking as laid down in this work.
“It is all rather poignant,” says Professor Philip Bejon, Director of the KEMRI-Wellcome Trust Research Programme and Dominic’s long-standing collaborator and friend. “The paper sets out a theoretical framework for how Dominic would spend the next chapter of his life. We've lost that. But what we do get is this very multidisciplinary, very scholarly piece of work, setting out all of the key issues and proposing various ways forward.”
Caroline Buckee first met Dominic during her PhD, describing her friend as always “generous and supportive in a mentorship role” to younger researchers. She is now Professor of Epidemiology at the Harvard T.H. Chan School of Public Health.
Professor Buckee also notes how fitting this paper is as his parting work. “It is a reminder of Dominic’s deep engagement with the scientific problems of the field. For his last paper to be this broad, visionary, deeply intellectual piece of work is remarkable.”
Grasping one of the thorniest problems in malaria research
The paper focuses on the transmission dynamics of malaria. Put simply: where do the mosquitoes that infect people get their malaria parasites from? This seemingly straightforward question has an incredibly complex set of factors to consider. As such, a failsafe method for working this out continues to elude researchers in malaria genomics.
Joining the Wellcome Sanger Institute’s malaria programme in 2007, Dr Bronwyn MacInnis worked side-by-side with Dominic as the global community around MalariaGEN took shape. “We were thinking about biology driving real-world impact from the very beginning. Dominic understood the potential, but also the very real challenges and risks of getting ahead of ourselves when trying to make statements about transmission dynamics based on genetic data. It wasn’t until he had enough data that he could go deep on this work.”

Mathematical models calculating transmission can be applied to other pathogens, such as viruses, and help public health authorities monitor and control diseases. But parasites like Plasmodium, which causes malaria, are more complex. As eukaryotic organisms, they reproduce sexually and thereby share genetic information from two parents, resulting in offspring with different sets of DNA than either parent. This is known as recombination.
The problem is confounded by the fact that people infected with malaria can have multiple lineages of the parasite in their blood at the same time. They might be infected in quick succession by mosquitoes carrying parasites with different genotypes (‘superinfection’), or bitten just once by a mosquito itself carrying multiple lineages of the parasite (‘cotransmission’). Throw in some human factors, such as migration, and the question of where the parasites in a person’s blood have come from becomes incredibly tricky.
“Recombination is this really thorny problem, and a defining problem of how we deal with the genomic epidemiology of malaria in the context of public health,” says Professor Buckee.
When trying to understand the spread of other pathogens, such as viruses, scientists typically construct a phylogenetic tree – something akin to a family tree connecting various virus lineages.
Dr MacInnis explains: “For other pathogens, you can build a tree and physically see this evolutionary information – how it relates to things that came before it. But for malaria you can’t really do that because the different pieces of the genome get mixed up through recombination, superinfection and cotransmission.
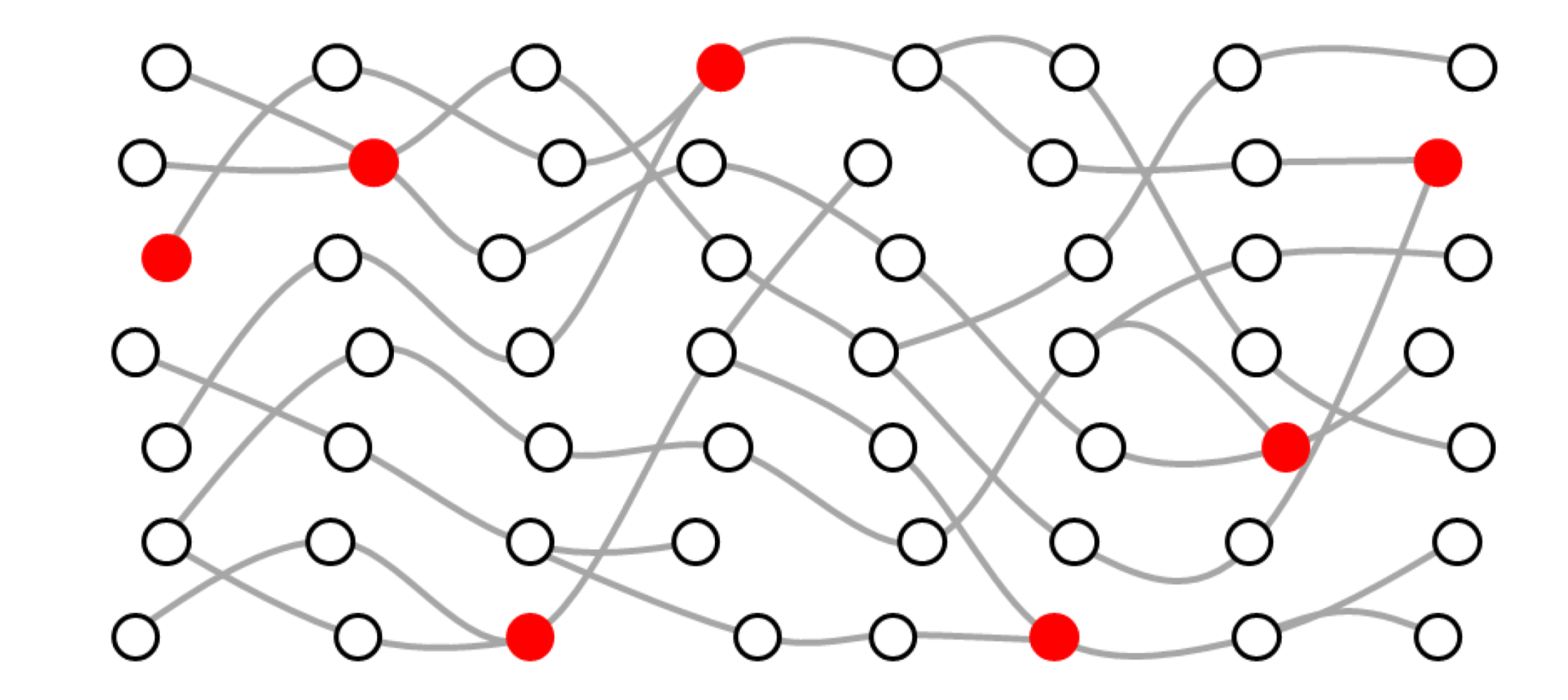
“Dominic was trying to find a visualisation that takes those factors into account but still creates a structure that allows you to connect those historical relationships in parasites.”
In his paper, Dominic proposes a genomic transmission graph. This shows the spread of parasites from host to host by mosquito vectors over time. It allows the different parasite lineages to be traced backwards and forwards in time while also clearly showing any potential recombination events within hosts.

Throughout the paper, Dominic dives deeply into what is happening at the points where parasites coalesce, proposing ways to calculate genetic variation of the parasites and potential population bottlenecks. He takes in the bigger picture, considering the interplay between global and local populations of parasites, as well as how to model changing epidemiological scenarios.
All this is backed up with a firm grounding in mathematics and bioinformatics. In fact, Dominic developed a new open-source coding package, a Python module called coalestr, to evaluate the genetic diversity of parasite populations.
Culmination of a career

“What the paper does is walk through a whole series of these different theoretical considerations that complicate things, dealing with a few interesting tangents on the way,” says Professor Bejon. “Any one of those could have been a paper in themselves. And so it's sort of multiple papers all concatenated together.
“There are not many people who could have written a paper like that. You need sight of both the biological and epidemiological aspects. You need some sight of how genomics works, particularly in the real world. And you also need a fair bit of mathematics. Dominic worked in each of those areas during his career.”
Much of what is being proposed in the paper would be impossible without the platforms, data sets, and tools that Dominic strove to build. Chief among those is MalariaGEN, an invaluable network of malaria researchers that Dominic was instrumental in setting up.
For Professor Buckee, Dominic’s efforts to build these resources are a large part of the reason that malaria research is as advanced as it is. “This paper gives a wonderful legacy of how he thinks about the analytical side of what we do with datasets like MalariaGEN. It is a reminder of Dominic’s deep engagement with the scientific problems of the field.
“He never lost his laser focus on the science. Despite MalariaGEN being this challenging consortium that he was managing – trying to grow it and support his collaborators – the scientific side, the methodological side, was always there in his head.”
A stepping stone for the community
This is not a science paper in search of headlines. It is written for the community Dominic surrounded himself with throughout his life, a global network of researchers dedicated to understanding the malaria parasite, its vector, and its host. And by studying the disease, he hoped it could be controlled and eliminated.
“I think this is one of those papers that we will come back to time and again, when we hit various blocks or start considering various problems, we'll refer back to this,” says Professor Bejon.
Professor Buckee agrees. “All my graduate students will have to read it. I think it outlines so much of the field. This is a deep, technical paper that is designed to be worked on and built on. It is part of the scientific stepping stones that we’re all on.”
Dr MacInnis is excited to see the reaction to the paper. “No doubt this will be a highly considered, highly digested piece of work that will open up new ways to think about the problem of malaria transmission dynamics. And as much as Dominic will have loved thinking through all these ideas and writing them down, what he really would have loved is putting it out in the world and engaging with other thinkers in our community.
“It’s such a shame he won’t be there to participate in that conversation. This is such a complicated problem that I’m sure there’s much more to the picture than even Dominic has figured out here. But this is a big step in a good direction. He will become the shoulders that future scientists in our field stand upon, taking this work in their own directions.”
Read the paper on Wellcome Open Research