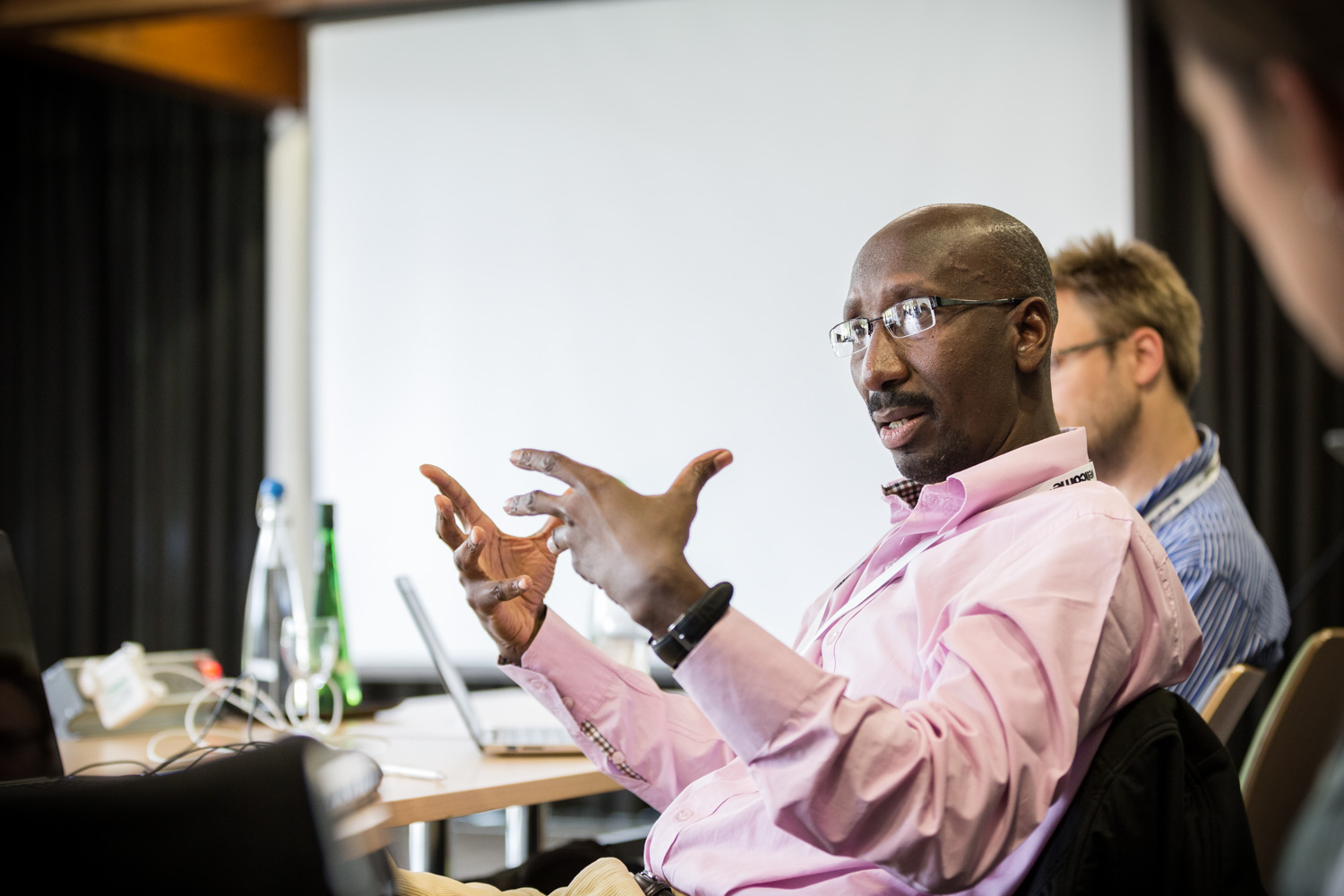
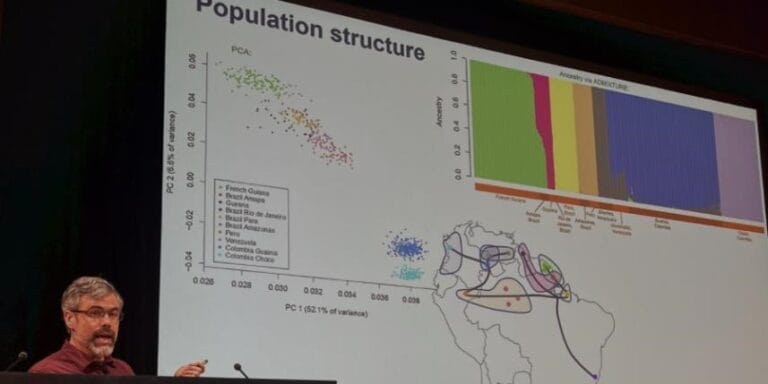
The annual Genomic Epidemiology of Malaria (GEM) meeting took place from 8-12 June at the Wellcome Genome Campus Conference Centre in Hinxton, UK. On a biannual basis, in the odd years, the meeting takes the form of a more focused, invitation-only retreat. In 2015, the overall aim of the meeting was to make progress with the analysis of large population genomic datasets for both the parasite and the vector, and to come up with concrete plans about how to use this type of data to build more accurate and responsive epidemiological models for guiding malaria control and elimination.
At this year’s gathering, we welcomed more than 50 scientists from 28 institutions. Participants formed three working groups – two driving forward existing projects, Pf3k and Ag1000G, and a third to consider ways of integrating population genetics into epidemiological modelling. These working groups engaged in highly-focused discussions and project-specific data analyses. Wider groups came together for joint working group sessions and plenary discussions. After each day’s intensive scientific work, participants had the chance to unwind and connect with a number of social activities including scientific speed dating, a bike ride and Frisbee.
See pictures from this year’s retreat.
Large-scale population genomics

Genomic epidemiology modelling
A major limitation of current epidemiological models is the lack of any detailed understanding of the natural diversity and demography of parasite and vector populations, of how parasite and vector populations evolve in response to interventions, and of what this can tell us about the likely success or failure of different control and elimination strategies. One of the most exciting challenges for malaria modelling over the next few years will be to unlock the vast amount of hidden information about parasite and vector populations that can be obtained by analysing spatiotemporal patterns of variation at high-resolution across the whole genome. This area was explored through joint sessions with the Pf3k and Ag1000G working groups, for example how to use large scale genome variation data to solve fundamental questions about spatial population structure, population dynamics and ongoing evolutionary processes, and how to capture all these interlocking processes in a coherent analytical framework.
See the GEM workshop programme for information on all workshop groups.
Acknowledgements
Many thanks to Matt Berriman, Martin Donnelly, Dominic Kwiatkowski, Mara Lawniczak, Gil McVean, Alistair Miles and Olivo Miotto who as members of the Scientific Organising Committee deserve much of the credit for the meeting. A big thank you also goes to the Wellcome Trust Scientific Conferences and Events Teams as well as to the MalariaGEN Resource Centre for all their hard work behind the scenes. And finally, a thank you to all participants who contributed so much to this highly fruitful meeting.