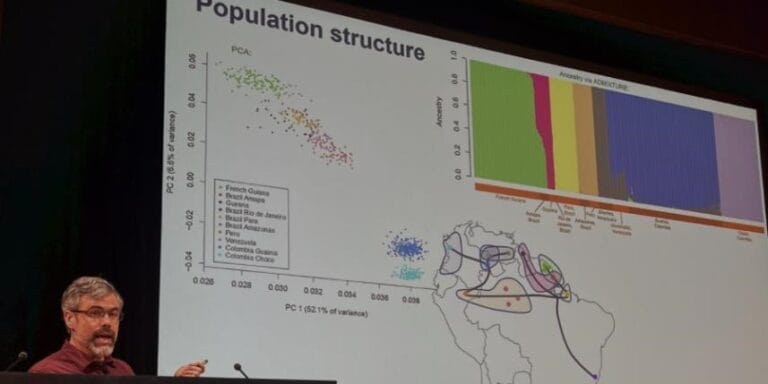
The recent release of 845 mosquito genomes by the Ag1000G Consortium provides the scientific community with a fantastic resource to investigate a huge variety of questions on a number of topics, such as speciation genomics, evidence for selection, and levels of population diversity.
Yet, working with such a huge dataset is complex. Just loading the genotype data into memory can be challenging – and that's before you even start on analysis!
We’d like as many people as possible to be able to explore and benefit from this resource, so this post provides a start-up guide to working with the Ag1000G data. In many cases you should be able to work on your local machine rather than a high memory cluster node.
File formats - VCF vs HDF5
VCF has been the de facto standard for working with sequence data for several years, particularly for single nucleotide polymorphism (SNP) data. VCF is a text-based format that is easy to understand by eye but computationally intensive to use, due to the large volume of file parsing involved.
Most workflows involve the reading in and parsing of a VCF file into computer memory where it is stored as a series of arrays. To make working with the data easier we provide an alternative data format where this hard work is already done: HDF5.
Using HDF5 it can be much easier, faster and more efficient to read data, e.g. genotypes, into memory for analysis. We extract data from VCF and store in HDF5 format using the vcfnp tool. The main advantage of HDF5 is that it can store data in binary format, and the data type (float/int/string) is defined in the file itself, so no expensive parsing is required.
Each HDF5 file can be thought of as a mini filesystem, with nested directories (aka groups) containing datasets. The structure used is described below, with the chromosome ID group at the root, containing two further groups (calldata and variants) and one dataset (samples).
Working with HDF5 files
HDF5 has implementations in R (h5; use v0.9.6 onwards) and python (h5py). The syntax of both is similar, here we load the first 10 rows of a dataset named foo located in the group bar2, itself within bar1:
python
import h5py
fh = h5py.File("path/to/file.h5", "r")
data = fh["/bar1/bar2/foo"][:10]
R
library(h5)
fh <- h5file("path/to/file.h5", "r")
data <- fh["/bar1/bar2/foo"][1:10]
Depending on how much data you load, this may take seconds to many minutes. We always need to be careful not to load too much data into memory at once.
Download HDF5 files from the Ag1000g phase 1 AR3 data release. For more information on these data, see the data release page.
More details of the format
A full description of HDF5 can be found here, but hopefully the following descriptions of the Ag1000G hdf5 files should get you up and running.
The group /{CHROMOSOME}/variants contains datasets that relate to variants. For example /2L/variants/POS describes positions of variants on chromosome 2L, /2L/variants/REF are the reference alleles of these variants and /2L/variants/ALT the alternate alleles. All fields that are in the INFO fields of a VCF can be found here. The 1st dimension of these datasets have size equal to the number of variants.
The group /{CHROMOSOME}/calldata contains datasets that relate to the variants and samples. Genotype data (/2L/calldata/genotype) is a 3-dimensional binary array; variants by samples by ploidy. Most other datasets are 2-dimensional, variants by samples.
The dataset /{CHROMOSOME}/samples contains the ID of each sample present. This is 1-dimensional, and is the size of the number of samples.
Importantly all data is position equivalent, so SNPs in calldata/genotype and in variants/POS have the same order, likewise, samples in /samples and columns of /genotype have the same order, so that it is easy to to filter out variants or samples based on a criterion and use the same filter to retain equivalent data.
Data analysis software
Python is our informatics team’s tool of choice for the analysis of Whole Genome Sequence (WGS) data, particularly with scikit-allel, a python module written by Alistair Miles.
Some worked examples in python can be found here. Additionally, have a look at Alistair’s blogs on Fst and PCA.
R does not quite have the support for dealing with compressed files in memory that Python does, so will tend to be less memory efficient, which may be problematic if you are working on a personal machine. Nonetheless, R has fairly comprehensive HDF5 support and excellent statistical and plotting tools.
Equivalent worked examples in R can be found here.
More from Nick on Twitter @nick_harding and GitHub https://github.com/hardingnj