The Bioinformatics Trainer Fellowship at PAMCA, in collaboration with the Genomic Surveillance Unit (GSU) at the Wellcome Sanger Institute, captivated my interest due to its unique combination of vector and malaria genomics. I became aware of the opportunity shortly after I had completed my MSc thesis, and it struck me that this was the best time to go for such an intense training in bioinformatics in order to contribute to providing solutions to current challenges in malaria vector control. I am fascinated by the potential of genomics to unravel the complexities of malaria transmission, vector biology, and host-parasite interactions. An opportunity to collaborate with esteemed researchers, engage in cutting-edge genomic projects, and play a pivotal role in training and mentoring aspiring bioinformaticians in vector and malaria genomics is incredibly exciting.
Through the fellowship program, I have been able to learn about a wide range of topics in vector control, all of which are geared towards enhancing the capacity for bioinformatics to support vector genomics work in Africa for malaria control and elimination. When I first joined the program, I had lots of questions about how things would turn out and the experiences that would come along with the training. It has only been a few months, but the learning has been so invaluable and fulfilling, and I am absolutely loving it. The fellowship involves rotations on different aspects of bioinformatics,including data analysis, data curation, variant calling pipelines, and scientific computing, all of which happen within the Vector Team of the GSU at the Wellcome Sanger Institute.
During the data analysis rotation (which I recently completed) my greatest highlight has been the personalised training. I got a chance to interact more closely with Dr. Alistair Miles, Dr. Christopher Clarkson, and the whole vector genomics surveillance team at Sanger. Both Chris and Alistair have a wealth of expertise in malaria vector genomics that they have been more than willing to share with us as fellows. Throughout the rotation, they not only offered guidance and great insights on how to navigate the data analysis tools, but have also shared their experiences on how they have managed to excel and thrive, something that I found very inspiring.
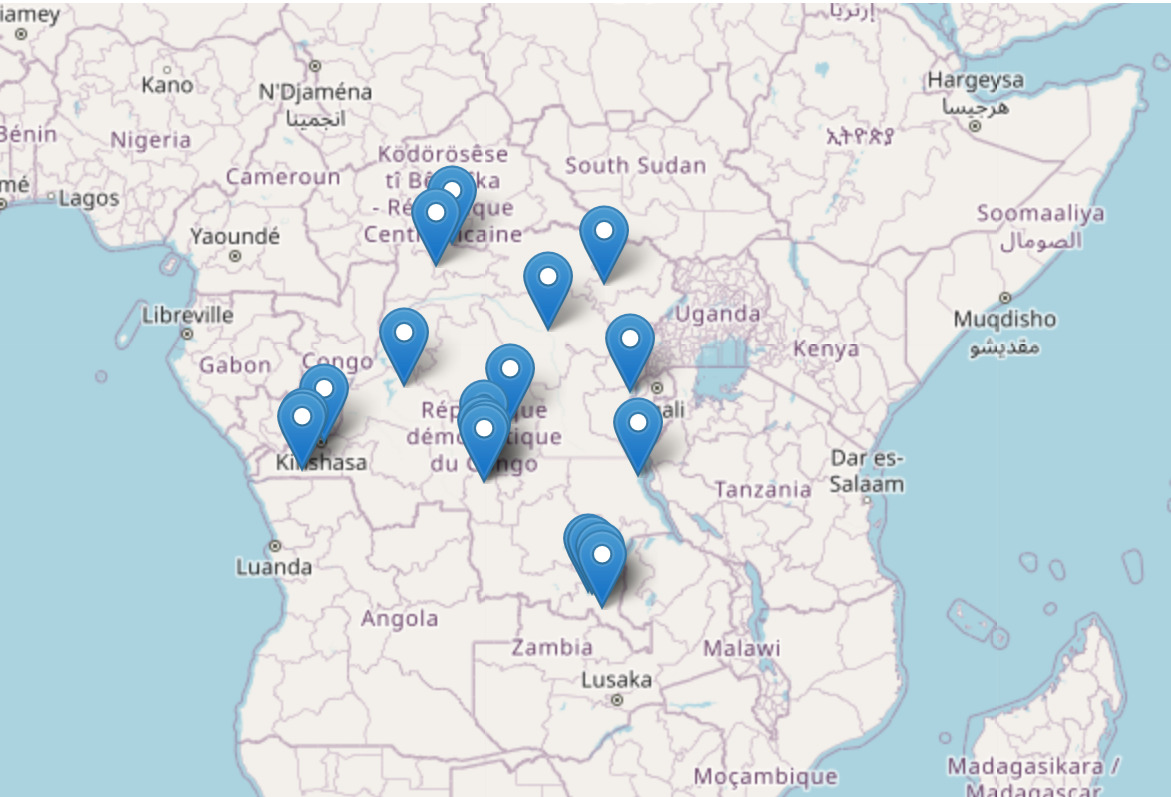
The data analysis rotation has also offered me an opportunity to have a feel of what real world challenges in malaria vector control are. For instance, I was able to work on the analysis of Democratic Republic of Congo sequenced Anopheles species, and generated useful insights on the trends of insecticide resistance by the malaria vectors, which is currently a huge threat to malaria vector control efforts. The ability to work with real data and generate new knowledge that can help answer questions on important topics such as population structure of malaria vectors, patterns and mechanisms of insecticide resistance, genetic diversity among sequenced mosquito populations, finding evidence of connectedness between the mosquitoes through adaptive gene flow, as well as identifying new regions under selection, have all made me appreciate more the potential of genomic tools in offering solutions to the current challenges in vector genomics.
Besides the great learning opportunity, the rotation on data analysis has also provided me with a platform to network with other professionals in different scientific fields, especially during discussions at the focus working groups, which is something I found very useful. Over the past couple of months, I have been able to learn and interact with various team members in a highly innovative environment and contribute to discussions more confidently. I have also had a chance to attend some interesting bioinformatics conferences; for instance, a recent one on Applied Bioinformatics and Public health Microbiology provided me with a deeper understanding of the genomics field. Having completed the data analysis rotation, I can confidently say that I have been able to gain a comprehensive understanding of the MalariaGEN data package, tools, and functions. These have been useful in generating essential outputs for making vector control decisions. What is even more important to me is that I got to do the actual analysis, building hands-on experience with a variety of software tools and techniques. It’s exciting to be able to apply my knowledge to real-world problems.
In general, being a Bioinformatics Trainer Fellow at PAMCA and the Sanger Institute has been extremely helpful in my career development as an early-career bioinformatics scientist. The fellowship provides exceptional training in malaria genomics and emphasises building the right skills and knowledge to contribute to malaria research. Through the first part of the fellowship, I have also been able to develop valuable interpersonal skills and improve on my ability to contribute more effectively to teamwork and collaborations. Without a doubt, the rotation has been very rewarding with lots of lessons learnt along the sides. I am excited about the fellowship, especially the fact that I get to challenge myself to learn new aspects in vector control.