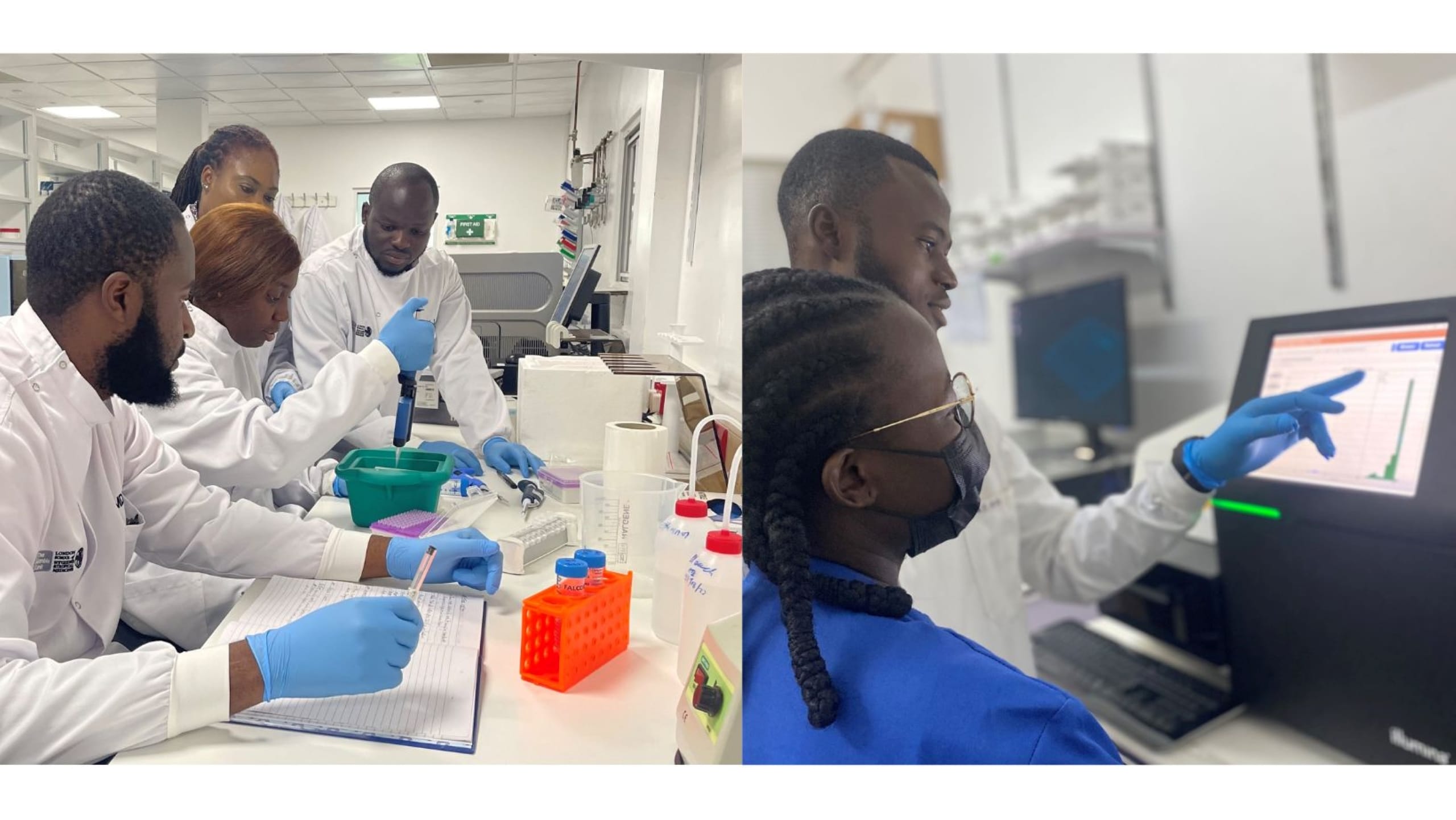
Whole genome sequencing (WGS) is a powerful tool for finding the genetic basis of insecticide and drug resistance. But it can be excessive, especially when the locations of relevant mutations are known.
Amplicon sequencing is a technique that selectively sequences only the parts of the genome where the mutations that we’re interested in occur. It’s faster and less resource-intensive than WGS. This makes it ideal for genomic surveillance.
MalariaGEN partners have been at the forefront of amplicon sequencing technology development and implementation. In collaboration with partners at sentinel sites in The Gambia, Ghana, Mali, Vietnam and Indonesia, we are developing open protocols for in-country amplicon sequencing. We are sharing the toolkit below for those wishing to create genomic surveillance networks using amplicon sequencing.
P. falciparum Amplicon Toolkit protocols
What’s inside
The MalariaGEN Amplicon Sequencing Toolkit provides partners with the checklists, protocols, and analytical methods needed to sequence and investigate targeted genes associated with drug or insecticide resistance. The amplicon process translates the raw sample data into accessible genetic report cards, and spatial-temporal maps of resistance, that can inform regional malaria control.
Until the DNA has been isolated and amplified, sample preparation and protocols differ for different species. Once the DNA is amplified, protocols for working with different species are the same.
For ease of use, we provide an end-to-end protocol for P. falciparum, P. vivax, and mosquitoes separately, which include reagent and equipment lists. For further information, please contact support@malariagen.net.