Malaria control efforts in South America face a growing and complex challenge as mosquito vectors continue to adapt to chemical control strategies. A recent study led by Jacob Tennessen and Dan Neafsey, at Harvard T.H. Chan School of Public Health, provides compelling new evidence that Anopheles darlingi, the primary malaria vector in the Americas, is evolving at the genomic level in ways that may reduce the effectiveness of insecticides commonly relied upon for vector control.
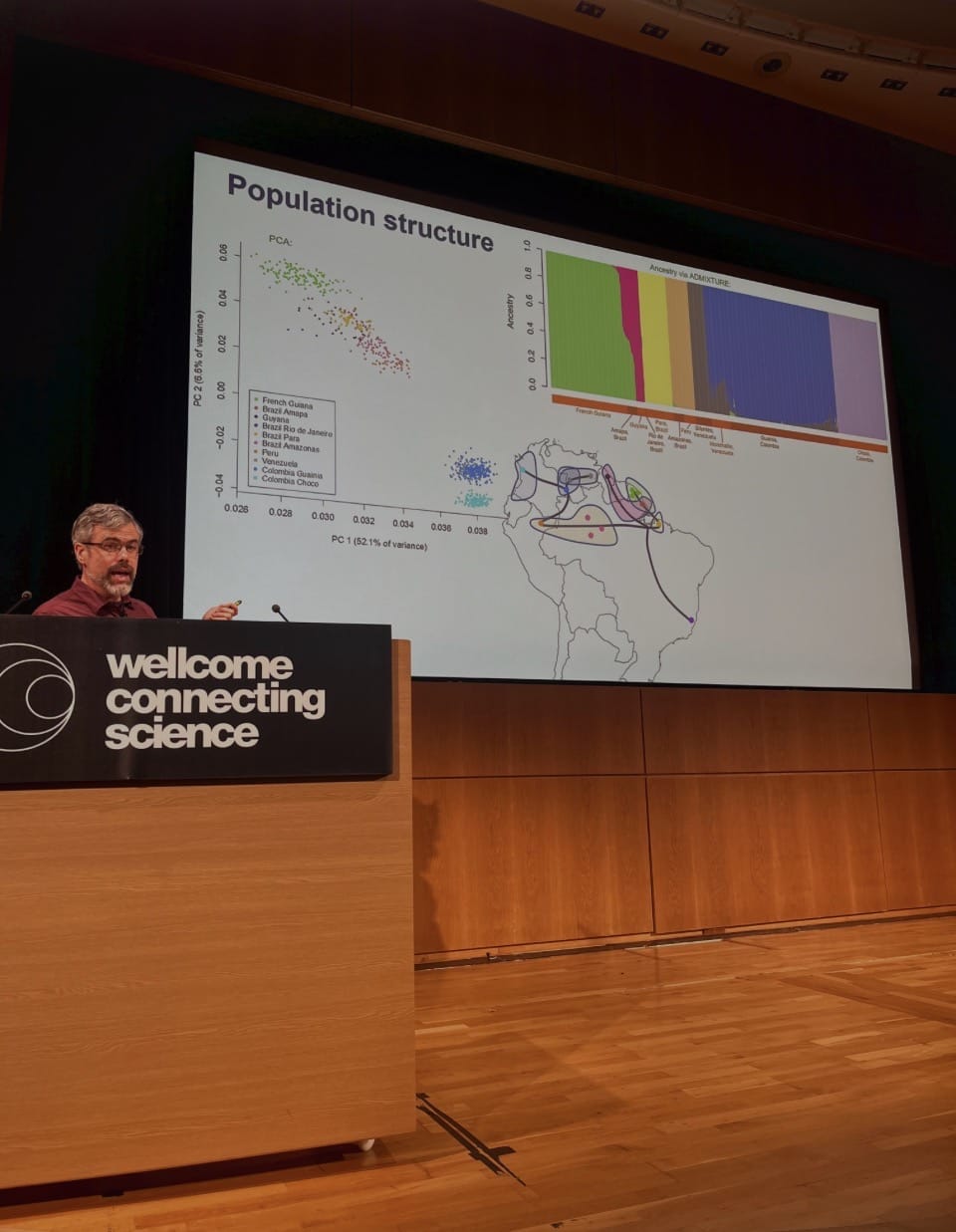
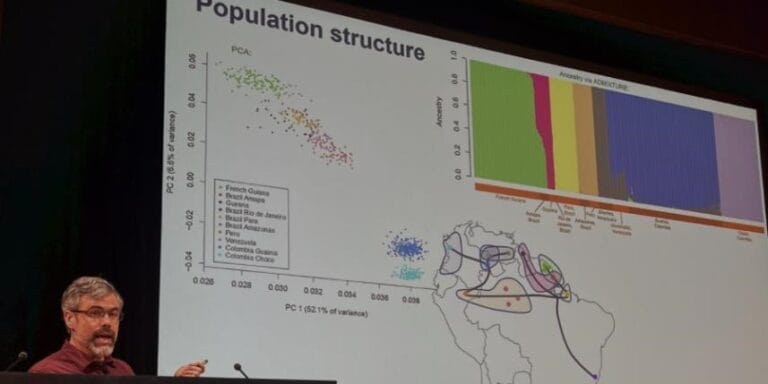
Published in Science in March 2026, the study represents the first large-scale whole genome sequencing effort of Anopheles mosquitoes in the region, analysing more than 1,000 genomes collected across six South American countries. The findings reveal widespread genetic signatures associated with metabolic insecticide resistance, detected across multiple populations and ecological settings. According to Neafsey, 'An association with agricultural settings suggests that public health control efforts may not be the driver of the observed genomic selection signals for metabolic resistance, but there may be phenotypic consequences for vector control nonetheless.'
While the study is framed as foundational research rather than an immediate policy intervention, its implications for malaria control are clear. Understanding how mosquito populations are evolving, where resistance-related adaptations are emerging, and how these changes are distributed geographically is essential for designing effective and sustainable vector control strategies. Whole genome sequencing is a powerful complement to entomological and epidemiological data already used by control programmes.
Importantly, the researchers have made sure that the value of this work extends beyond South America and reinforces the importance of open, well-curated data resources that enable comparative analyses across settings and over time.
These themes resonated strongly during discussions at a previous Genomic Epidemiology of Malaria (GEM) conference, where the idea of hosting the Anopheles darlingi genomic data resource within MalariaGEN, as part of the Vector Observatory, was first explored. The Vector Observatory is designed to support large-scale, standardised and accessible genomic surveillance of malaria vectors, facilitating collaboration and knowledge exchange across regions.
Reflecting on those discussions, a Dr Vikki Simpson noted: “We are delighted to have the privilege of hosting this incredible resource. It is a testament to the collegiality of Jacob, Dan, and their teams, whose commitment to open science and collaboration has made this possible.”
As genomic data increasingly inform public health practice, partnerships built on trust, shared infrastructure and collective stewardship will be critical. Hosting the An. darlingi resource within MalariaGEN’s Vector Observatory represents more than data sharing. It reflects a shared vision for how genomic insights can be mobilised across regions to strengthen malaria control and preparedness in the face of evolving vector populations.
By connecting cutting-edge research with global surveillance platforms, this work helps ensure that discoveries made in one region can inform action worldwide, reinforcing genomics as a cornerstone of modern malaria control.