Drug resistance once again threatens progress against malaria. Beginning in 2008, clinical studies in Cambodia reported that the frontline antimalarial drug, artemisinin, was taking longer to clear the parasites from the blood of malaria-infected patients—a worrying sign that drug resistance was emerging, again (1-3).
At the time, artemisinin was used in combination with a partner drug, piperaquine, which helped to keep the treatment effective. The combination of dihydroartemisinin and piperaquine (DHA-PPQ) seemed to contain the threat, but then in 2013, researchers began reporting a new outbreak of multidrug resistance—this time to DHA-PPQ (4). The simultaneous failure of both drugs renders the combination therapy powerless to treat these multidrug resistant infections. The situation is fragile, and doctors are left to rely on a small number of drugs.
By now this latest threat is well-documented (5-9), and not entirely surprising. Malaria parasites have evolved to evade each and every antimalarial drug that we’ve used to treat them–chloroquine, sulfadoxine pyrimethamine, mefloquine, you name it.
Biologically speaking, it’s only a matter of time. Each new drug creates the pre-conditions for resistance: a selective pressure. The more we use the drugs, the more selective pressure we apply. Parasites with a genetic make-up that happens to make them better able to survive the drug are more likely to survive long enough to reproduce—gifting this ability to the next generation and increasing the frequency or proportion of their genes.
Biological fitness—in this context, how successful parasites are at passing on their genes—is somewhat easier to measure in reverse. “If there are a great many descendants alive today, that tells us something about the biological fitness of their ancestors, but it doesn’t tell us the whole story of how they came to be so successful,” says Dr Roberto Amato, Senior Staff Scientist at the Wellcome Sanger Institute.
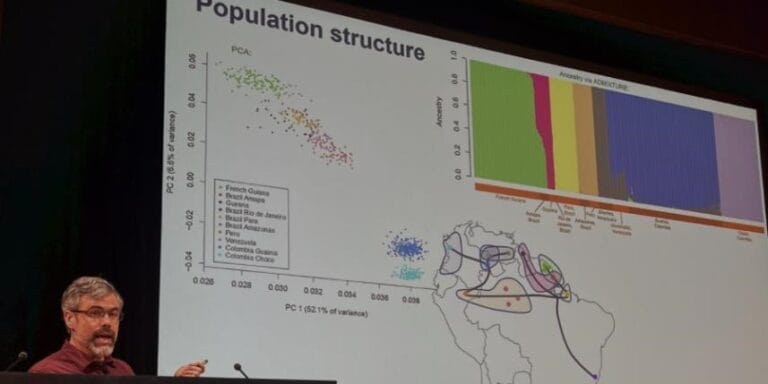
“We had the data to look back in time and observe the rise of multidrug resistance in the region,” he adds, referencing an analysis published this week in Lancet Infectious Diseases. Amato and his colleagues compared the genomes of nearly 1,500 P. falciparum samples collected from malaria patients in 11 locations in Southeast Asia between 2007 and 2013—the years spanning the adoption of DHA-PPQ as a frontline treatment to the first clinical reports of resistance.
As it turns out, by the time DHA-PPQ drug resistance was detected in the clinics of western Cambodia, the evolutionary battle had been raging for years. Amato and colleagues showed that mutations associated with drug resistance were already circulating five years before the first clinical studies documented the problem.
Identifying drug resistance as soon as possible is only one (albeit important) advantage of undertaking genetic studies of drug resistance—it can also offer important insights into the complex evolutionary process.
For example, two parasite samples can have the same mutation yet not be related to each other. Disentangling these relationships gets at the heart of some very important uncertainties that surround an outbreak of drug resistance, for example, whether it is spreading or emerging independently in different locations.
A good example of this is artemisinin resistance, which is associated with genetic changes in a gene called kelch13.
“Malaria parasites in western Cambodia developed artemisinin resistance rapidly and repeatedly. We found more than 30 distinct lineages—like branches on a family tree, these lineages track related parasites. Each lineage is characterized by a particular kelch13 mutation with strong similarities in the surrounding DNA sequence, and represents a possible distinct emergence of artemisinin resistance. Six of these lineages contain the infamous 580Y kelch13 mutation. Yet only one of them, KEL1, became dominant,” explains Amato.
That wasn’t the end of the story. In 2008, the KEL1 artemisinin-resistant lineage combined with PLA1, the major lineage associated with piperaquine resistance.
“More than 90% of the DHA-PPQ resistant parasite samples in our study belonged to the KEL1/PLA1 co-lineage that emerged in the very same year that DHA-PPQ became the first-line antimalarial drug—and we can see that it’s spread aggressively, displacing other artemisinin-resistant strains,” warns Amato.
It’s clear that drug resistance is a perennial problem. With large, high-quality data sets, genetic and genomic studies can be used for regular monitoring of drug resistance, and to inform periodic responses—working to manage resistance in a population, much the way a physician might manage chronic illness in a patient.
MalariaGEN’s SpotMalaria project is working to integrate genetic surveillance into the routine monitoring done by national malaria control programs (NMCPs). And, methodologically the foundations are there: a single drop of blood collected on a piece of filter paper contains enough parasite DNA to test for genetic markers of drug resistance. This technology is cost-effective, reliable, and easy to implement in the field.
“There’s an opportunity to systematically monitor genetic changes in parasite populations for early signs of drug resistance and track its spread. In turn, this allows us to control the balance of interventions, preserve the useful life of existing drugs, and help to ensure effective treatment,” says Amato.
This will be all the more crucial in mounting a coordinated and effective response in times of crisis.
References
- Noedl H, Se Y, Schaecher K, Smith BL, Socheat D, Fukuda MM. Evidence of Artemisinin-Resistant Malaria in Western Cambodia. N Engl J Med 2008; 359: 2619–20.
- Dondorp AM, Nosten F, Yi P, et al. Artemisinin Resistance in Plasmodium falciparum Malaria. N Engl J Med 2009; 361: 455–67.
- Ashley EA, Dhorda M, Fairhurst RM, et al. Spread of Artemisinin Resistance in Plasmodium falciparum Malaria. N Engl J Med 2014; 371: 411–23.
- Leang R, Barrette A, Bouth DM, et al. Efficacy of dihydroartemisinin-piperaquine for treatment of uncomplicated plasmodium falciparum and plasmodium vivax in Cambodia, 2008 to 2010. Antimicrob Agents Chemother 2013; 57: 818–26.
- Saunders DL, Vanachayangkul P, Lon C. Dihydroartemisinin–Piperaquine Failure in Cambodia. N Engl J Med 2014; 371: 484–5.
- Amaratunga C, Lim P, Suon S, et al. Dihydroartemisinin-piperaquine resistance in Plasmodium falciparum malaria in Cambodia: a multisite prospective cohort study. Lancet Infect Dis 2016; 16: 357–62.
- Imwong M, Suwannasin K, Kunasol C, et al. The spread of artemisinin-resistant Plasmodium falciparum in the Greater Mekong subregion: a molecular epidemiology observational study. Lancet Infect Dis 2017; 17: 491–7.
- Thanh NV, Thuy-Nhien N, Tuyen NTK, et al. Rapid decline in the susceptibility of Plasmodium falciparum to dihydroartemisinin–piperaquine in the south of Vietnam. Malar J 2017; 16: 27.
- Imwong M, Hien TT, Thuy-Nhien NT, Dondorp AM, White NJ. Spread of a single multidrug resistant malaria parasite lineage ( PfPailin ) to Vietnam. Lancet Infect Dis 2017; 17: 1022–3.