This open access resource provides a foundation for further research into how genetic variation and sexual recombination affects parasite biology, at a much higher resolution than previously possible. These data are being made available at a time of intense interest in studying the genetic basis for evolutionary changes in the malaria-causing P. falciparum parasite, such as the emergence and spread of antimalarial drug resistance.
“A lot of progress has been made in recent years in mapping out variation in the P. falciparum genome, however there are still big gaps in our knowledge, including many genes that are relevant to vaccine development or drug resistance,” explains Alistair Miles, Head of Informatics with the MRC Centre for Genomics and Global Health. “These new data on the parasite crosses provide a solid foundation for extending our knowledge into these clinically important regions of the genome.”
The P. falciparum crosses have each previously led to landmark discoveries. The first cross proved that parasites undergo sexual recombination whilst inside the mosquito. The second cross revealed the gene responsible for chloroquine resistance. The third cross led to discoveries about the way in which parasites invade human blood cells. All three crosses continue to play an central role as model systems for studying parasite genome biology, and this data resource will enhance their value.
Each parasite cross is analogous to a large family, with two parents that mate to produce dozens of offspring. Like human families, each offspring has a genetic make-up inherited from its parents via a process of recombination which reshuffles the parental DNA sequences. This sexual recombination is central to how parasites evolve and adapt to new pressures such as antimalarial drugs, and the new P. falciparum crosses data resource provides a unique opportunity to study parasite recombination in detail.
The P. falciparum crosses can also help to unlock information in genome regions that have proved difficult to study. Although modern DNA sequencing technology has transformed the study of genetic variation in malaria parasites, analysing the raw sequence data is a complex process with many sources of error. The crosses provide a unique means to cope with this complexity. Each offspring should inherit its DNA from one of its two parents. If an offspring appears to have a genetic make-up that is not found in either parent, that suggests an error in the data. These inconsistencies can be used to learn about different types of error, and design strategies to remove them.
“Because the crosses give us this guidance on which data to trust, we’ve been able to gain new insights about different types of genetic variation, including insertions and deletions, complex sequence changes, and copy number variation,” says Miles. “We hope the data are a useful resource for the research community, and we welcome feedback on how the resource could be developed and improved.”
This data release is accompanied by a preprint which highlights several interesting features of the data, and is being made available to the wider scientific community on the preprint server, bioRxiv.
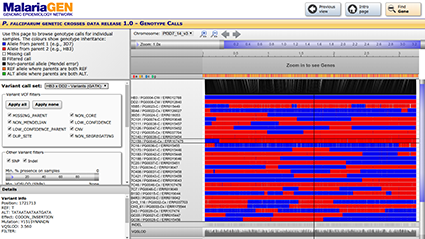
The P. falciparum crosses project data release 1.0 is available open access.
To learn more and to download data, visit: https://www.malariagen.net/data_package/pf-crosses-1-0/
Explore the data online: https://www.malariagen.net/apps/pf-crosses
Preprint details
Miles A et al, 2015. Deep sequencing of Plasmodium falciparum genetic crosses: a resource for the study of genome variation and meiotic recombination. bioRxiv doi: http://dx.doi.org/10.1101/024182