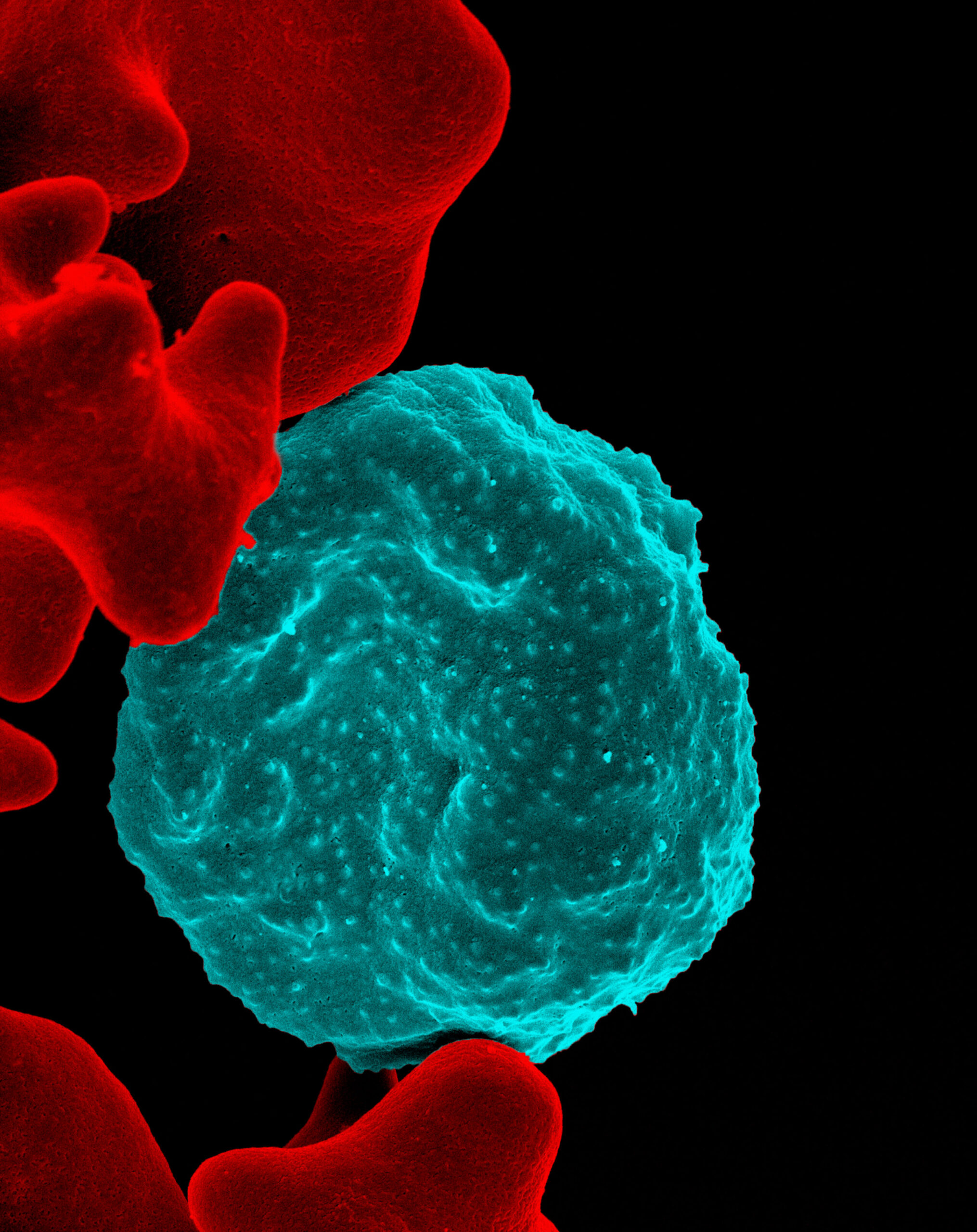
Malaria-infected red blood cell. Photo credit: NIAID, Flickr 2013, CC-BY2.0.
To hunt for possible molecular markers, researchers started with an African strain of P. falciparum parasites that had been repeatedly exposed to artemisinin until it was able to survive high doses of the drug in the lab. By comparing the whole genome sequence of the lab-cultured resistant strain with a sibling strain that had not been exposed to the drug, they were able to identify several mutations of interest in a number of genes.
The team then excluded from consideration those mutations that weren't also present in lab-cultured parasites collected in Cambodia, where artemisinin resistance first emerged. This further narrowed their target to a number of mutations in the K13-propeller that were strongly associated with increased parasite survival rates when exposed to artemisinin in the lab.
The team then investigated whether the distribution K13-propeller mutations correlates with artemisinin resistance in Cambodia by sequencing the K13-propeller domain of malaria parasites collected in several provinces in Cambodia between 2001-2012. They found that in provinces where artemisinin resistance is known to exist, the frequency of K-13 propeller mutations increased significantly over time — an increase not observed in areas without artemisinin resistance.
To determine whether K13-propeller mutations are a useful molecular marker in clinical settings, the team studied K13-propeller mutations in patients from the Cambodian provinces of Pursat and Ratanakiri and for whom they knew the parasite clearance half-lives. These samples had previously been sequenced and genotyped through the MalariaGEN P. falciparum Community Project and assigned to artemisinin-sensitive or artemisinin-resistant subpopulations in the analysis described in Miotto et al (2013). The researchers found that patients who carried parasites with K-13 propeller mutations took significantly longer to clear malaria parasites from their blood than patients carrying parasites without these mutations.
Taken together, these findings suggest that mutations in the K13-propeller play an important role in determining artemisinin resistance, act as a molecular marker for artemisinin-resistant parasites, and provide important clues into how resistance is developing in parasite populations in Southeast Asia.
Publications
Ariey et al. A molecular marker of artimesinin-resistant Plasmodium falciparum malaria. Nature. 2014 Jan 2;505(7481):50-5. doi: 10.1038/nature12876. Epub 2013 Dec 18.
To read more about the effort that led to the discovery and the importance of these findings, see:
Plowe. Malaria: Resistance nailed. Nature. 2014 Jan 2;505(7481):30-1. doi: 10.1038/nature12845. Epub 2013 Dec 18.